Table of Contents
How to
This is a collection of short answers and links to some miscellaneous questions that we have had while doing research in Oncinfo. This can also be a useful resource for other scholars in the field of computational biology with similar interests and challenges.
Measure memory used by GPUs?
Use torch.cuda.memory , e.g., to discover the effect of clearing gradients at the end of each iteration. With ignite, this can be done using an event handler that calls optimizer.zero_grad(set_to_none=True).
Analyze multiomicds data?
Neurepiomics workshop in 2023:
- Shiva's presentation on omicsdatabases
- Mohsen's presentation for Neurepiomics workshop, 2023-10-09.
- Habil's slides on multiomics data integration.
What is the best Linux distribution for biologists?
ArchLinux and its derivatives, like Manjaro, are the top Linux distributions for biologists. The BioArchLinux community maintains an excellent repository with over 4,000 bioinformatics packages and their dependencies. Regular automated updates ensure the latest versions are available by fetching source code from trusted repositories and compiling releases optimized for the target architecture. Its philosophy is described here.
Download files to where you cannot login or click using a GUI?
Login using GUI on another computer. Click to start downloading. While it is downloading, right-click on the page and then click on “Inspect”. Then, click on “Network ” on the top of the panel and find the file. Get the cURL command and run it in terminal to download the file in the other computer.
Supercharge your YouTube experience and minimize ads?
“Enhancer for YouTube” is a popular browser extension that enhances the YouTube viewing experience by providing additional features and customization options, but it may contains malware! While it doesn't explicitly focus on blocking ads, it offers some options to minimize or hide them. Use Autoskip to automatically skip ads that pass the Enhancer filter.
The above solutions reduces the income of the person or the team who created and posted the YouTube video. Better than the above solutions is to use a YouTube premium account. In this way, you a) save your time and focus by avoiding ads and b) support the person who created the video. Youtube pays creators a little money for every minute we watch their video with a premium account.
Prevent Google Colab from being disconnected?
To prevent disconnection of your Google Colab session, you can use the following script that keeps the session active by clicking on the notebook every 60 seconds:
function ClickConnect(){
console.log("Working");
document.querySelector("colab-connect-button").click()
}
setInterval(ClickConnect,1000*60)
To implement this script, open the developer console in your browser (using Ctrl+Shift+J in Chrome or Ctrl+Shift+I in Firefox or Command+Option+C in Safari), and paste the above script in the console. This will help to ensure that your Colab session remains active and does not disconnect due to inactivity.
Plot deep neural network architecture diagrams?
There are several apps that would help in this manner which are listed based on their descending qualities:
- PlotNeuralNet (very simple, just generate fixed latex commands, no flexibility in the layers themselves, you need to rewrite models in this language)
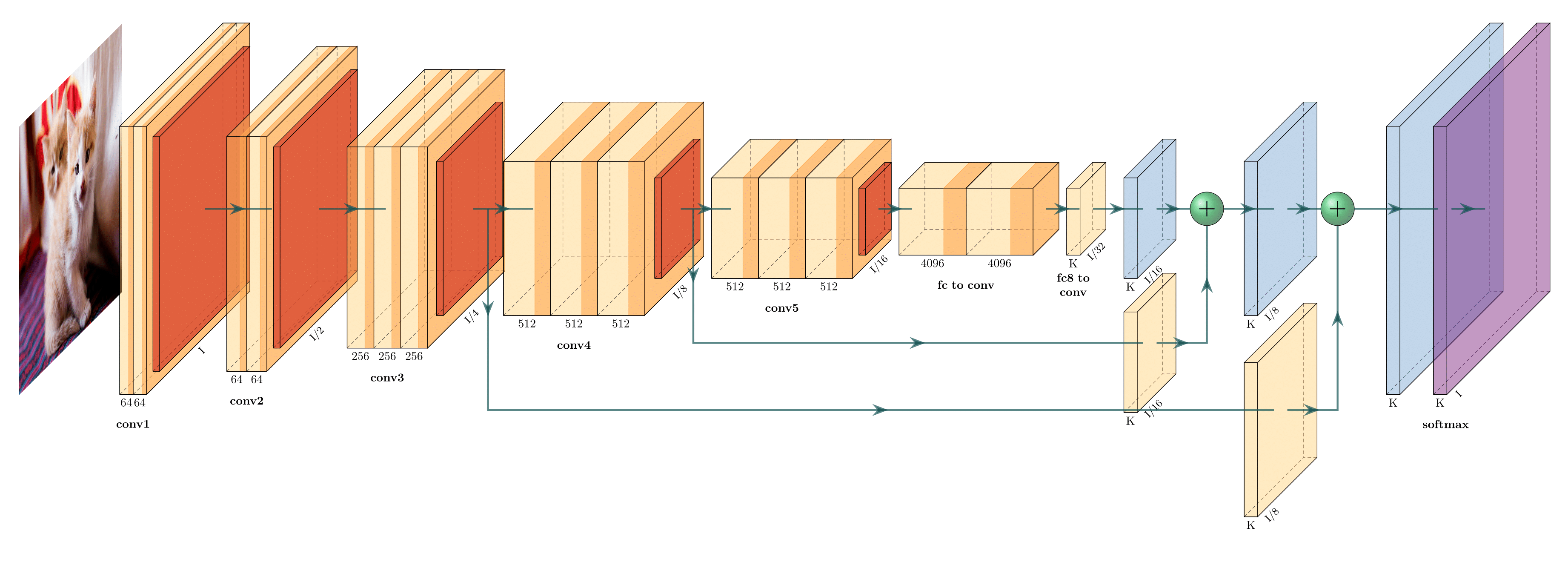
- Visualkeras (plots Keras models)
- netron (does not consider ModuleList inside the model in PyTorch)
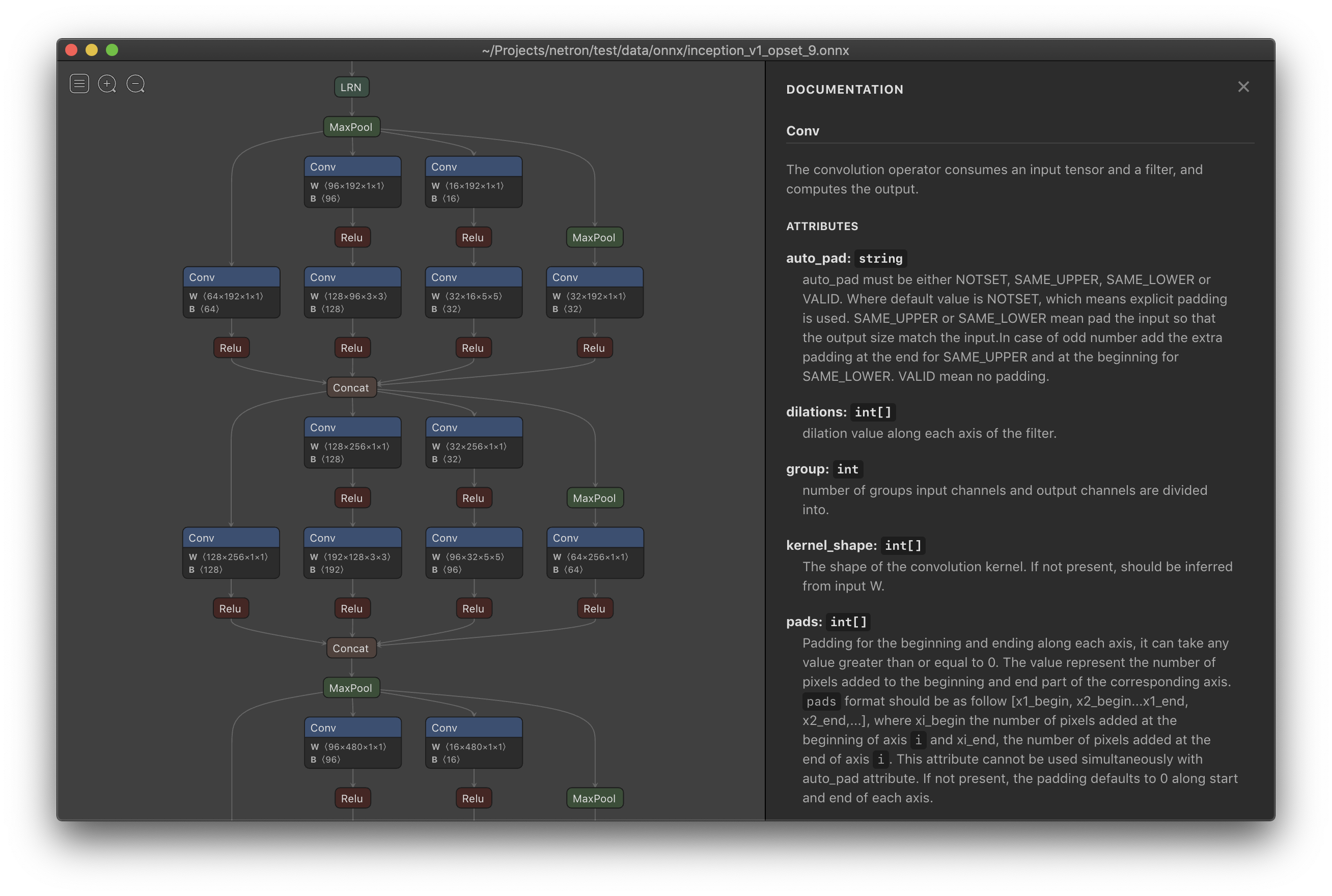
- Net2Vis (automatically generates abstract visualizations for convolutional neural networks from Keras code)
- LaTeX examples of neural networks.
Append multiple videos quickly?
First, make a text file named mylist.txt with the following content:
file part1.mp4
file part2.mp4
file part3.mp4
Then, run
ffmpeg -f concat -safe 0 -i mylist.txt -c copy output.mp4
which is fast because it only copies the audio and video codecs. ffmpeg can be installed on MacOS using brew.
Extend reads in a bam file to a fixed fragment size to better view peaks with IGV?
Assume you have input.bam, which contains mapped short single-end sequence reads, but you know the fragment size=100. With the following command, you can make a bam file that has the extended reads, but you will miss the actual reads. This helps to better visualize the actual read signal:
bedtools bamtobed -i input.bam | \
awk -F'\t' 'BEGIN {OFS = FS} {if ($6=="+") {$3=$2+99} else {$2=$3-99}; if ($2<0) {$2=0}; print $0;}' | \
bedtools bedtobam -g hg19.chr.sizes -i -> extended_input.bam
where hg19.chr.sizes is a tsv file with two columns containing chr names and sizes.
NOTE: bedCoverage command is supposed to do this, but in my case, I got an error about the input bam file, despite the fact that it was a normal mapped bam file.
Run simultaneous jobs (on multiple files) in a single mpi on a cluster?
It can be done using xargs -P <number-of-threads> <command>. For example, we assume you are writing a bash script as follows:
First, write a function (or script) to do the work on a single input file. Second, export the function using
export -f my_func_for_single_file
Third, use the `find` command (or anything appropriate) and pipe the results to `xargs` like this:
find . -type f -name "*.bam" -printf "%f\n" | xargs -P 125 -I {} bash -c 'my_func_for_single_file "$@"' _ {}
Where %f is the basename of the file returned by the find . This command runs the task with 125 threads in parallel on a node (e.g. with 128 cores). Assuming you have 210 files, It will run all the task in 2 sweeps, which means it costs less almost with a factor of 125.
Change 125 as you want, but not more than the number of available cores in each compute node! :) Also you can adapt this idea to the cases where your function needs other inputs than the simple file name.
Plot models of Keras in R on MacOS?
First, in Terminal , run this brew command (which is much better than “port”):
brew install graphviz
Then run this in R:
reticulate::py_install("pydot", pip = TRUE)
Identify genes of a pathway in R?
1) ReactomeContentService4R is Reactome wrapper in R:
library(ReactomeContentService4R) event2Ids(event.id = "R-HSA-6790901") # can only input id here
2) Kegg:
library(org.Hs.eg.db)
kegg <- org.Hs.egPATH2EG
mapped <- mappedkeys(kegg)
kegg2 <- as.list(kegg[mapped])
keggFind("pathway", "mtor") ## finds the pathway with a key word
ent <- kegg2 [["04150"|]] ##needs id of pathway (mtor), returns entrez id
gene.mapping(id=ent, inputType="ENTREZID", outputType = "SYMBOL")
Work with screen sessions?
There are five main commands while working with screen session:
- Start and name a screen:
screen -S <NAME of the screen> - Detach from a screen:
Ctrl+a d - See the list of active screens:
screen -ls - Reattach to a screen:
screen -r <NAME of the screen> - Quit and kill your screen:
Ctrl+athenCtrl+\
Read and write excel files in R
Use openxlsx package to read, write and edit xlsx files in R. Package's integration with C++ makes it faster and easier to use. Simplifies the creation of Excel .xlsx files by providing a high level interface to writing, styling and editing worksheets. Through the use of 'Rcpp', read/write times are comparable to the 'xlsx' and 'XLConnect' packages with the added benefit of removing the dependency on Java.
E.g. Writing four dataframes in four sheets of excel workbook can be done as follows:
library(openxlsx)
data("mtcars")
listDataFrames <- list("First"=mtcars[1:5,], "Second"=mtcars[10:15, ])
xlsFile <- "~/temp/test.xlsx"
w1 <- write.xlsx(x=listDataFrames, file=xlsFile)
## You can add new sheets:
addWorksheet(w1, sheetName = "New")
writeData(wb=w1, sheet="New", mtcars[,1:3])
saveWorkbook(w1, file=xlsFile, overwrite=TRUE)
## Read a sheet in a data frame:
r1 <- read.xlsx(xlsxFile=xlsFile, sheet="Second")
Set local mirror for Rscript
Put following lines above your script.
local({r <- getOption("repos")
r["CRAN"] <- "your mirror, example: http://cran.us.r-project.org "
options(repos=r)
})
Add only modified changes and ignore untracked files using git?
Using newer version of `git` (e.g., >=2.32.1),
git commit -am 'minor changes'; git push
⚠️ Caution: With older versions the -a option will add all files in the directory, which is usually NOT what you want. Instead, use following command, which would have the same effect as above:
git ls-files --modified | xargs git add; git commit -m 'minor changes'; git push
The above should work if there is no conflict. If there is a conflict at push time, first pull. Now, you need to look for “»>” in the code, and manually fix the conflict. Then, push again.
Create an R package or use a package that is being developed?
An R package source folder has a predetermined structure. If you change the content of a packge, you can use the make.package() function to update (i.e., build, check, and install) the package while preserving its structure like the senescence and iNETgrate package examples.
Cancel all your jobs on the Stampede?
squeue -l -u $USER|awk 'FNR>2 {print $1}' |xargs scancel
Install R locally (e.g. on a cluster)?
Like most of Unix programs, R can be installed from source by a) downloading the source code, configuring and compiling the code, and then installing the binaries. If you try this simple approach and your get errors like these, it means the dependencies are not available or updated on your machine. E.g., f you want to install the latest development version of R on your macOS, first install Fortran if you do not have it. You may also need to update PCRE using brew on macOS. Alternatively, you can compile PCRE2 from the source, and then let R where it is using CPPFLAGS and LDFLAGS.
If you do not have sudo permissions, like when you are working on a cluster, you should either use the software module, or install R locally in your home directory. E.g., you can install R on Stampede or Maverick TACC clusters as follows:
mkdir ~/tempR; cd ~/tempR ## Make a temp directory wget http://cran.cnr.berkeley.edu/src/base/R-3/R-3.2.2.tar.gz ## Always get the latest version tar -zxvf ./R-3.2.2.tar.gz; cd ./R-3.2.2 ## Unzip, the rest is by following INSTALL file that is in this folder. mkdir ~/arch ## You will install here. ./configure --prefix $HOME/arch make make check make install
If the above works without any error, you can add $HOME/arch to your path by inserting the following line in your .bachrc file:
export PATH=$HOME/arch/bin:$PATH
On some clusters, a few libraries might not be installed or they might be too old (e.g., zlib, curl, bzip2, xz, pcre). In particular, the bzip2 issue can be resolved by following these steps on the Lonestar5 cluster. Oncinfo Lab members can use it if they add the following to their .bashrc
export PATH=/home1/03270/zare/Install/bin:$PATH
Installing R using conda is only a quick and dirty, temporary solution. E.g., as of 2021-05-14, the xml2 package that is installed by conda is not compatible with R 4.0 that is installed using conda, therefore, solving the issue in this way moves the R version to from 4.0 back to 3.0! The time you will spend addressing such issues would be possibly more than the time you need to put on to a clean instalation of R from source.
Restore a file deleted in a local git directory?
Use git reset –hard to completely bring your working directory to HEAD state. However, this is a dangerous command because you may loose some local files that are not pushed yet.
Get familiar with machine learning and its applications in computational biology?
- You can enroll in many online machine learning courses. Some of the best courses in ML can be found here.
- Most common ML techniques are very well explained in Scikit learn with illustrations and example Python code. These techniques have been implemented in R packages including mlr3 and tidymodels.
-This 2015 paper reviews applications of ML in genetics and genomics. Read it and follow its references.
Get access to the papers through the library when you are off-campus?
In any of these two ways:
a) First add the following to your browser (e.g., Chrome, Safari, or Firefox) bookmarks.
 For your convenience, you can write “javascript”, and then “:voind()”, and copy the following line between the parentheses.
For your convenience, you can write “javascript”, and then “:voind()”, and copy the following line between the parentheses.
location.href=%22http://libproxy.uthscsa.edu/login?url=%22+location.href
Then, on the journal page, click on the bookmark. Login and start reading.
b) Use GlobalProtect, which is the University VPN.
If an articles is not available from the library, it can be ordered via interlibrary loan.
Convert pdf to MS word?
Try whatever you can to avoid conversion! Instead, educate your team and your collaborators to use Authorea, Overleaf or at least Google Doc. In Google Doc, references can be easily handled using Paperpile add-on (NOT the extension), and figures can be automatically numbered using the the Cross Reference add-on as suggested in these guidelines on how to write academic documents with Google Docs. Add-ons are not available when editing .docx files. Only if your biologist collaborators cannot unfortunately edit the LaTeX source, consider using a conversion tool such as Adobe Acrobat Chrome extension or Acrobat Pro, which can export a .pdf as a .doc file. docs. The docs zone and pdf Converter are online alternatives. If you need to to separate pages, use pdfjam. If Bibtex is not an option, use EasyBib.
Docx2LaTeX can convert simple Word or Google Docs to LaTeX.
Enable spell check in Emacs on OS X?
The default Aquamacs spell checker has some issues. To replace it, first install Aspell, which is a replacement for Ispell:
brew install aspell
And then add the following line to your emacs initialization file, e.g., ~/Library/Preferences/Aquamacs Emacs/Preferences.el, or ~/.emacs
(setq ispell-program-name "/usr/local/bin/aspell")
Do microaray analysis or anything else in Bioconductor?
This is an excellent site with many well commented code examples and a lot of handy short-cuts. See also Functional analysis tools.
Learn to learn?
Take this online course: Learning How to Learn: Powerful mental tools to help you master tough subjects.
Improve your writing?
Read and write a lot. Have someone proofread your write up. Use tips and resources that can help you improve your writing skills to the next level. E.g., you can find valuable professional resources on Writing Forward website. Coursera has a special course on writing emails in its “Improve Your English Communication Skills Specialization”.
Prepare a scientific poster?
Decide on the main figures that you want to include. Put them in a template and write a caption for each figure. Write your abstract and an appropriate title. Send it to collaborators and ask for their comments at least a week before printing.
Find a biological database of interest?
First, look at the list of biological databases. Also, if you know an important database that is missed in this list, please add it as a service to the community.
Begin learning bioinformatics?
Take a course from the list of free online bioinformatics courses e.g., the Computational Molecular Biology Course at Stanford is broad and covers the classic topics but it is not updated, and may become outdated. The same is true for PLOS Translational Bioinformatics Collection of articles, which are more advanced. Most central topics are covered in some course from the European Bioinformatics Institute (EBI). Very useful training materials are available from GOBLET. Videos from the Models, Inference & Algorithms Initiative (MIA) at Broad are relatively advanced.
Bioconductor course material provides a walkthrough in specific areas. Also, you can find valuable resources from the list of bioinformatics workshops and conferences. Following the “Bioinformatics for biologists workshop” does not require expertise in computer programing. Bio-Linux team prepared a good introduction, which can be used as a reference too. If machine learning is new to you, read about its applications in genetics and genomics. You need to have some basic knowledge in mathematics and statistics too.
Decide on appropriate courses for a program in computational biology?
Browse the Online Computational Biology Curriculum, which lists hundreds of courses available from Coursera, Edx, etc, and comments on their relevance to computational biology. Berkeley and Stanford also have lists of relevant courses.
Establish and maintain a computational biology lab?
Do not miss About My Lab, a valuable collection of PLOS articles on how to manage a lab. This is a fundamentally difficult job.
Install Salmon on OSX?
If you do not have autoconf, install it. Following the installation guidelines, for OSX you need to first install Thread Building Blocks (TBB) (brew install tbb) and then check that the installation was successful (brew list). Download the latest version of Salmon source code and uncompress it. Follow Salmon's installation guidelines. The cmake command in the guidelines will be something like the following for OSX:
cmake -DFETCH_BOOST=TRUE -DTBB_INSTALL_DIR=/usr/local/lib -DCMAKE_INSTALL_PREFIX=/usr/local/
Follow the rest of the guidelines. You may need to use (sudo make install) if you get a permission error.
Write a scientific paper?
Put the figures together and then draft different sections. Focus the Discussion. Be careful about authorship. It might be easier to write the abstract after other sections are drafted. You can use your own voice. Explain in the cover letter how your work would be interesting to the journal audiance.
Many journals require vectorized figures and different tweaks to the fonts and colors. So, make sure your figures are easily reproducible. At the submission stage, you can change to a vectorized format like eps or pdf using different tools including Inkscape.
Prepare or review computational biology papers for Nature methods?
Read their “Reviewing computational methods” (2015) and “Guidelines for algorithms and software in Nature Methods” (2014) articles. Provide source code, pseudocode, compiled executables, and the mathematical description. Softwares must be accompanied with documentation, sample data and the expected output, and a license (e.g., GPL≥2). Have a look at The list of computational biology papers in Nature Methods published in 2015, and the hints by an editor of Nature Communications. Several journals publish papers in this field.
Set the default width of fill mode (line length) in emacs?
Use 'M-x customize-variable' to set 'fill-column' (100 in Oncinfo). Use DejaVu Sans Mono (~Menlo on MacOS) size 18-20 is an appropriate font for programming in Emacs. To do so, you may need to manually edit your .emacs in macOS, and add the following line:
(setq truncate-lines nil)
Get older versions using git?
Use “git log” to see the previous commits and the corresponding hashes, “git checkout <hash>” to get an older version, and “git checkout master” to get back.
Learn about linear models and ANOVA in R?
Review Advanced Statistical Methods II lecture notes by Dr. Larry Ammann at UT Dallas.
Convert gene or protein IDs?
bioDBnet, BioMart - Ensembl, or AnnotationDbi package in R to convert between Entrez Gene, RefSeq, Ensemble, and many more.
Prepare attractive, scientific presentations ?
Use a “home slide”. Also, learn about other tips from Susan McConnell.
Access a Bioconductor package source code?
It is always better to a install the latest version of a package as directed in the corresponding Bioconductor page (e.g., Pigengene). If you need to see more details in the source code, or you need the development version. If the package maintainer adds your public ssh key, then you can clone the source from the Bioconductor using the “Source Repository (Developer Access) ” command, which is posted on the corresponding package page, e.g.,
mkdir ~/proj; cd ~/proj git clone git@git.bioconductor.org:packages/Pigengene
Now, you can build the package fom the source using:
R CMD REMOVE Pigengene; R CMD build Pigengene
If the build is successful, a tarbal will be created. You can install the new package using:
R CMD INSTALL Pigengene_<Version>.tar.gz
Use git via proxy or vpn?
Use sshuttle, e.g., sshuttle -r h_mailto:z14@nyx.cs.txstate.edu 0.0.0.0/0 -vv
The list of servers at Texas Sate University are listed here.
Enable autocomplete and the tab bar in Emacs?
Install the auto-complete and tabbar packages from MELPA. In OS X, you may need to edit your init file (usually it is .emacs but you can use C-H v user-init-file RET to check) file to remove auto-complete from the package-selected-packages list and instead add the following lines:
(require 'auto-complete) (global-auto-complete-mode t)
Also, read the packages instructions to learn how to configure and use them.
Update 2024-08-14: tabbar can take 50%-90% CPU usage! Use tab-bar-mode in Emacs >=27.
Cite references?
Find the original paper that is most relevant. Setup recommendation engines not to miss recently published work.
Resubmit an NIH grant?
Address ALL reviewers' comments. Write a strong introduction [ppt ]. Watch NIH Webinars for Applicants.
Silence a gene?
Small interfering (si) RNAs and miRNAs bind to mRAN and prevent it from being translated.
Compare the data resources on a gene across databases?
Xena provides heatmaps for one gene using databases selected from a list of tens of databases from TCGA,1000 genomes, etc. Cancer Genome Cloud allows the user to integrate their own tools with theplatform and run it on, say, TCGA in the cloud. Seeks orders thousands of datasets based on how concordant the expression of the input genes are.
Save Powerpoint in high resolution on OS X?
Select all, open Preview > File > New from Clipboard. DPI is different from PPI.
Download TCGA normal data?
Use “Add cases filter” link to add sample_type as a filter.
Know about the immune system?
It is a prerequisite (2012) to understand immunotherapy (2015). CAR-T Cells (2014) are engineered outside the body to express receptors specific to a patient's particular cancer. Immune checkpoint (2017) inhibitors can release the break of the immune system.
Use computational biology in immunotherapy?
A) Mine the gene expression or mutation databases to discover tumor antigens (targets of immune system). B) Combine these data with mass spectrometric data from tumor specimens, or sera, to identify immunogenic (actionable) antigens. C) Analyze gene expression profiles to predict which patients will benefit (also, have benefitted) from a specific treatment (e.g., infer infiltration of immune cells in tumors). D) “Evaluation of humoral response against a diverse set of self-antigens can be explored rapidly using the high-content protein or peptide microarrays” (Thakurta et al.). E) Assess the diversity of the vast 10^14 possibilities for T-cell receptors using proteomics or sequencing. Many tools have been developed to address these needs (2017 review).
Discover immunogenic antigens (neoantigens) suitable for immunotherapy?
Identify the somatic mutations in expressed genes, predict epitope in silico using publicly available databases, and immunize mice with long peptides encoding the mutated epitopes to determine immunogenicity.
Avoid misinterpretation of biological experiments?
Reasoning must be logical. Report enough details of the methods to reproduce the results. Assess the robustness of the findings with respect to minor perturbations to the experimental settings. To prove that drug A targets protein X, it is not sufficient to confirm that treatment with A leads to killing cells that have X. Maybe the cells are killed because of some other mechanism. Use “rescue experiments” as in the A=imatinib X=BCR–ABL case. Always, avoid these ten common statistical mistakes.
Analyze single cell RNA-seq data?
Map the reads similar to bulk RNA-seq data, exclude outlier cells and genes, cluster the cells, and project the cells onto an annotated reference such as Human Cell Atlas. The broken links can be found in the Sanger's course.
Make interactive 3D plots in R?
Use plot3D::scatter3D to create the graph:
scatter3D(x=iris$Sepal.Length, y=iris$Petal.Length, z=iris$Sepal.Width, clab = c("Sepal", "Width (cm)"))
plotrgl()
and then use plot3Drgl::plotrgl to interactively zoom and rotate it.
Improve the ranking of your institution better than MIT?!
Hire scholars who are frequently cited as adjunct professors.
Rescue US biomedical research from its systemic flaws?
Hypercompetition has damaging effects. Reduce the number of PhD students and postdocs. Revise funding mechanisms to encourage “path-breaking ideas that, by definition,” are risky. Support early-stage investigators.
Test an R function that you have extracted from a script?
Sometimes you have a working script but some part of it is useful also elsewhere. You extract that portion and make a function f(x,y) out of it. Now, in the script you call that function. You run your modified script and the output is exactly the same as before. Is this a sufficient test for the function you just wrote? No! What if you forget to include one of the variables as an input argument of the function? Then, that variable, which is still defined somewhere in the script before you call the function, is used in the function as a “global variable” without any explicit error. However, when the function is used in a different context, that global variable may not be defined, or worse, it may have an irrelevant value. To avoid this issue, use codetools::findGlobals(…, merge=FALSE) to identify and remove all global variables from your function. A cumbersome and less accurate way to identify global variables follows:
- Put a browser in the script right before calling the function.
- Save the function and its inputs in a temporary file, e.g., save(x,y, f, file=“~/temp/f1.RData”)
- Remove all objects in this R session using rm(list=ls()).
- load(“~/temp/f1.RData”).
- Type “c” and “Enter” to run the rest of the script. The output must be exactly the same.
- If the function uses a global variable, while running it this way, you will probably get an error like “object 'z' not found”. Then, either compute z in f, add it as an input argument of f, or modify the line that uses z.
- Repeat until you get no errors, and then, check that the output is exactly the same as before.
Separate the reads corresponding to each individual cell from a single-cell RNA-Seq fastq file?
Use the barcodes and Unique Molecular Identifiers (UMIs).
Copy from your prior clipboard history?
Use a clipboard manager like Clipy (developed based on ClipMenu) or JumpCut. Clipy allows you to save lists of favorite or frequent text snippets for later use.
Review a paper?
Read the journal guidelines, and the Nature's quick and concise tutorial. The PLOS Open Reviewer Gateway can be helpful if you are interested in becoming a reviewer for PLOS journals. Read BMC or Nature introduction, and take the Nature's Focus on Peer Review course.
Get credit for your reviews via Web of Science Reviewer Recognition Service (publons).
Adjust the acceleration and speed of a Logitec mouse?
Install and use the Logitech Control Center.
Encrypt a folder?
Encrypt a largeFolder folder using tar and compress it using gpg based on the AES-256 encrypting algorithm. You will obtain the strongest security with these options:
tar -cvz largeFolder | gpg --s2k-mode 3 --s2k-count 65011712 --s2k-digest-algo SHA512 --s2k-cipher-algo AES256 --symmetric --no-symkey-cache -o largeFolder.tgz.gpg
The –no-symkey-cache option is available in version >=2.2.7. On macOS, you need to first install GnuPG. An alternative approach is to use 7z, which can be installed using homebrew, however, 7z is windows based and thus not recommended.
To decrypt and uncomperess,
gpg --decrypt --no-symkey-cache largeFolder.tgz.gpg | tar -xv
tar may need -z –no-same-owner.
A good password should have at least 12 characters, include both small and capital letters, and at least one digit and one special character such as !@#$%^&*(). Do not use dictionary words in your password, instead, use a passphrase “to create strong passwords”.
Upload a file to Oncinfo and link to it?
Set the link type to “internal media”, click on “Browse Server”, and then on “Choose File” and “Update”.
Aks people's opinion?
Create a Doodle to find a common time for scheduling events. For longer surveys, use SurveyMonkey for offline, and Poll Everywhere for online interaction with audience.
Disable scroll acceleration in macOS?
Install and use USB Overdrive to set Wheel up and down “Speed” of your mouse to say, 6 lines. The following command does NOT work:
defaults write .GlobalPreferences com.apple.scrollwheel.scaling -1
Logitech Control Center may help on the Logitech MX mice older than 2019.
Choose a solid state (SSD) external drive?
The non-volatile memory express (NVMe) devices are better than SATA solid state drives. Good brands include Sabrent (Nano is smaller than Pro but gets hot when extensivly used), Seagete, Addlink, and Team. As of 2020, a speed of 1000 Mb/s is possible using USB 3.2.
Search for public domain images?
Unsplash is among the best resources with many hi-resolution images, which are frequently used by media.
Identify Senescence in Cells and Tissues?
Watch this quick 6-minute introduction to the senescence concept, and learn about the common markers and kits.
